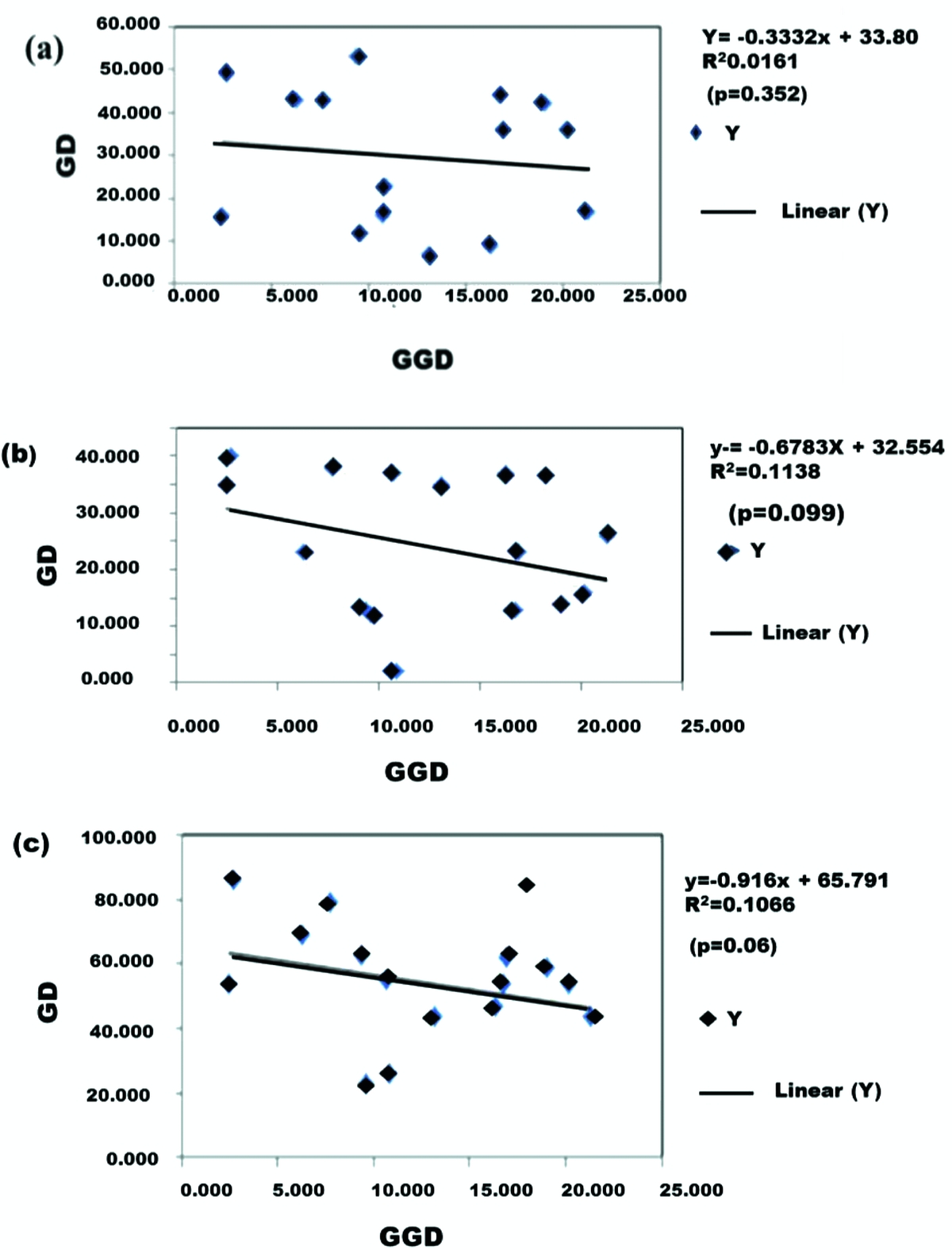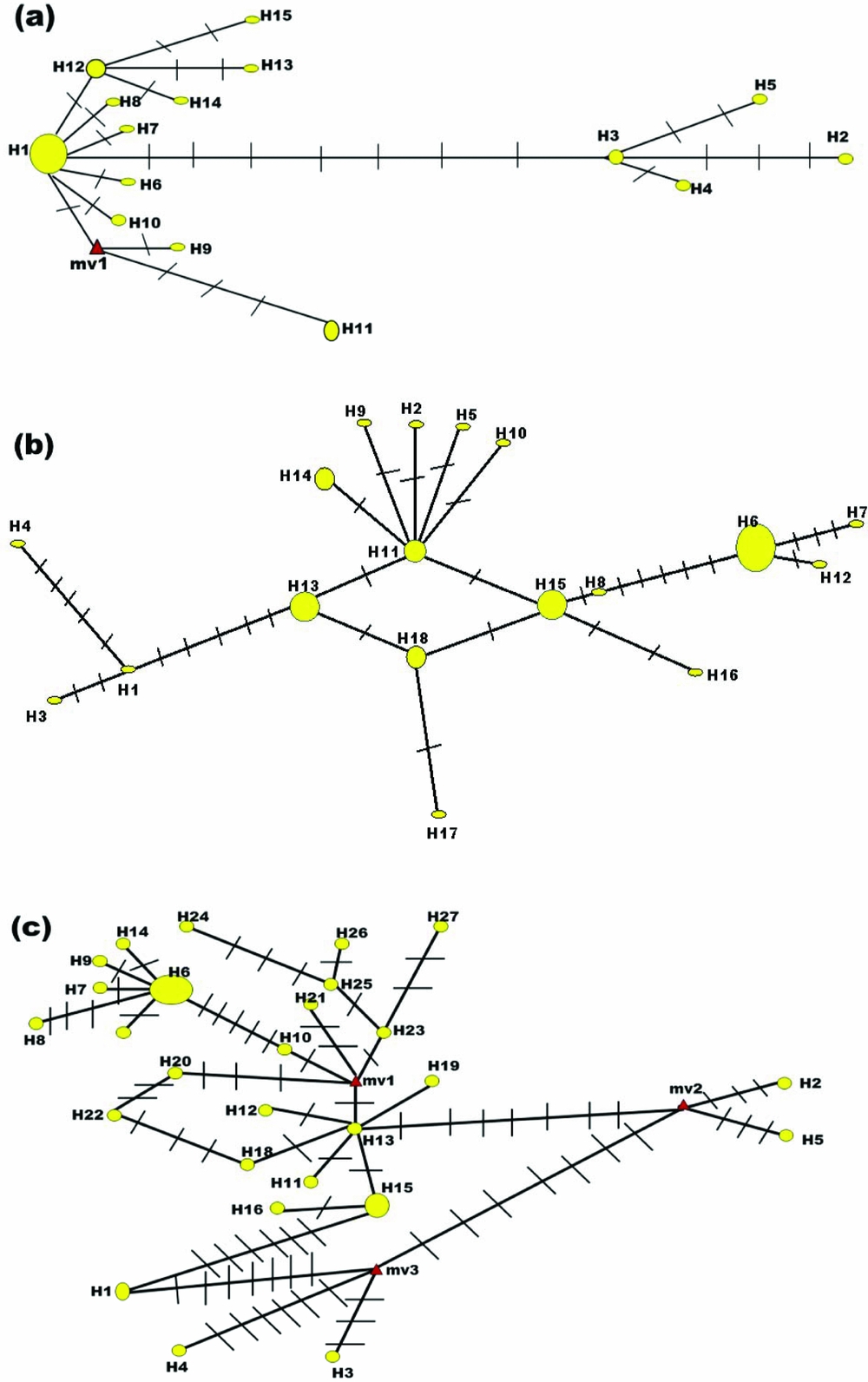
Analyses of the internal transcribed rDNA spacers (ITS1 and ITS2) of Indian weevils of Odoiporus longicollis (Olivier) reveal gene flow between locations | International Journal of Tropical Insect Science | Cambridge Core

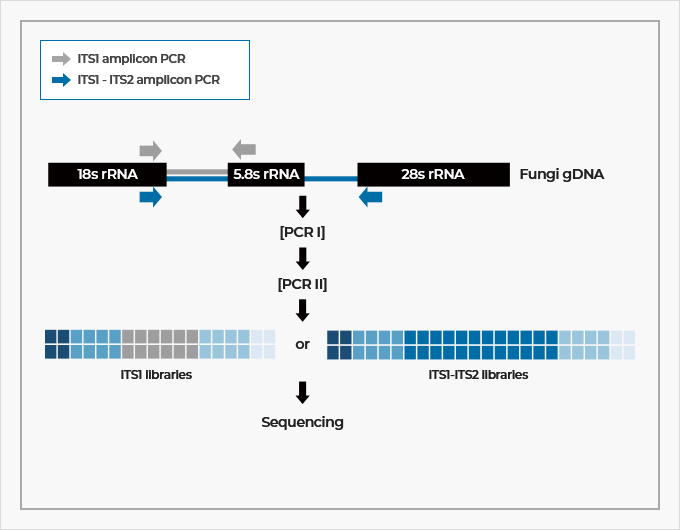
JCI Insight - Customization of a DADA2-based pipeline for fungal internal transcribed spacer 1 (ITS1) amplicon data sets

General Structure of a Nuclear Ribosomal RNA Gene ITS1 and ITS2 are... | Download Scientific Diagram
![Comparison of the performance of ITS1 and ITS2 as barcodes in amplicon-based sequencing of bioaerosols [PeerJ] Comparison of the performance of ITS1 and ITS2 as barcodes in amplicon-based sequencing of bioaerosols [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/8523/1/fig-3-2x.jpg)
Comparison of the performance of ITS1 and ITS2 as barcodes in amplicon-based sequencing of bioaerosols [PeerJ]
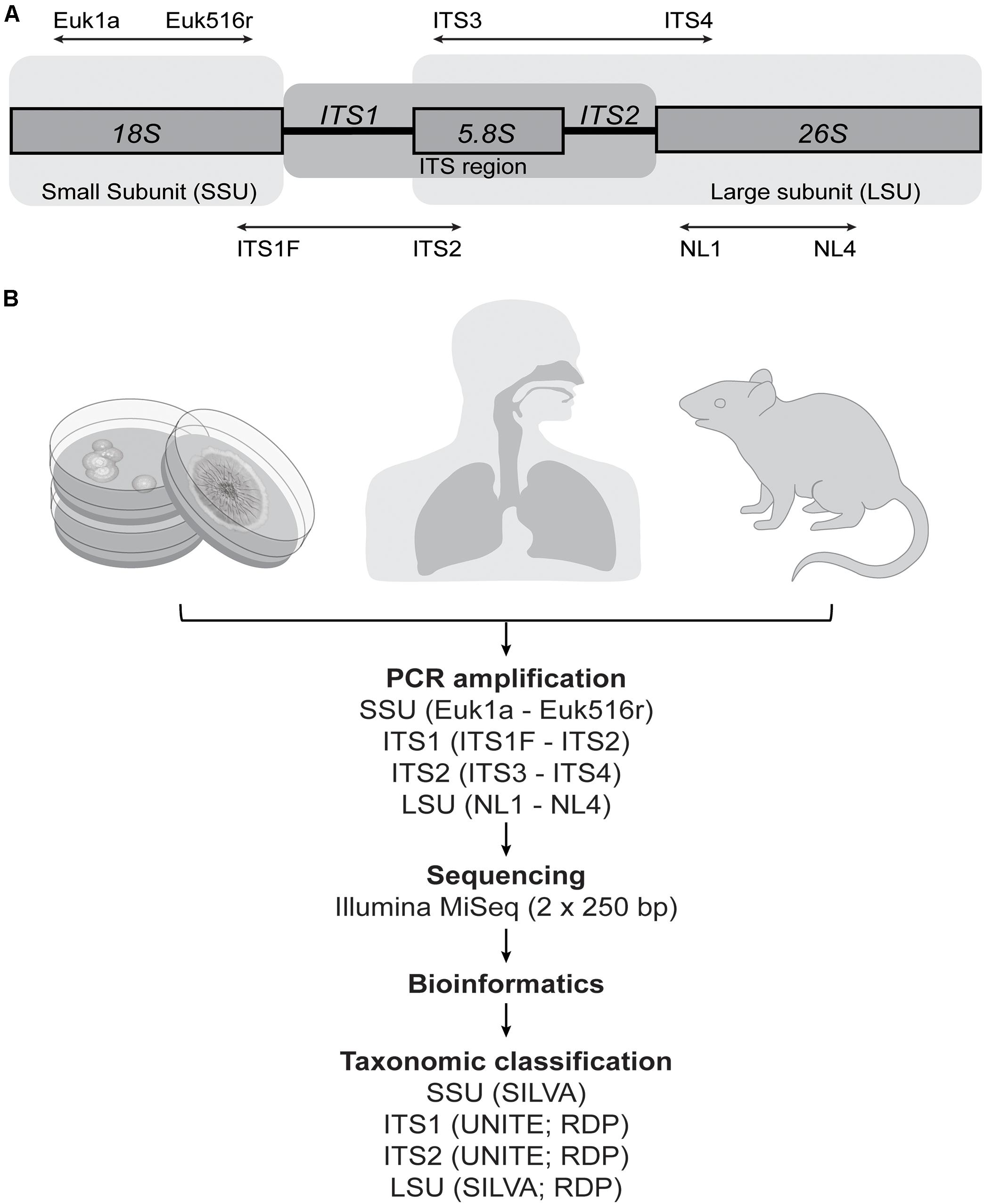
Frontiers | Characterizing the Human Mycobiota: A Comparison of Small Subunit rRNA, ITS1, ITS2, and Large Subunit rRNA Genomic Targets
ITS1 Library Preparation Kit for Illumina (Cat. 71000, 71010, 71020, 71030, 71040) | Norgen Biotek Corp.
Evaluation of the ribosomal DNA internal transcribed spacer (ITS), specifically ITS1 and ITS2, for the analysis of fungal diversity by deep sequencing | PLOS ONE
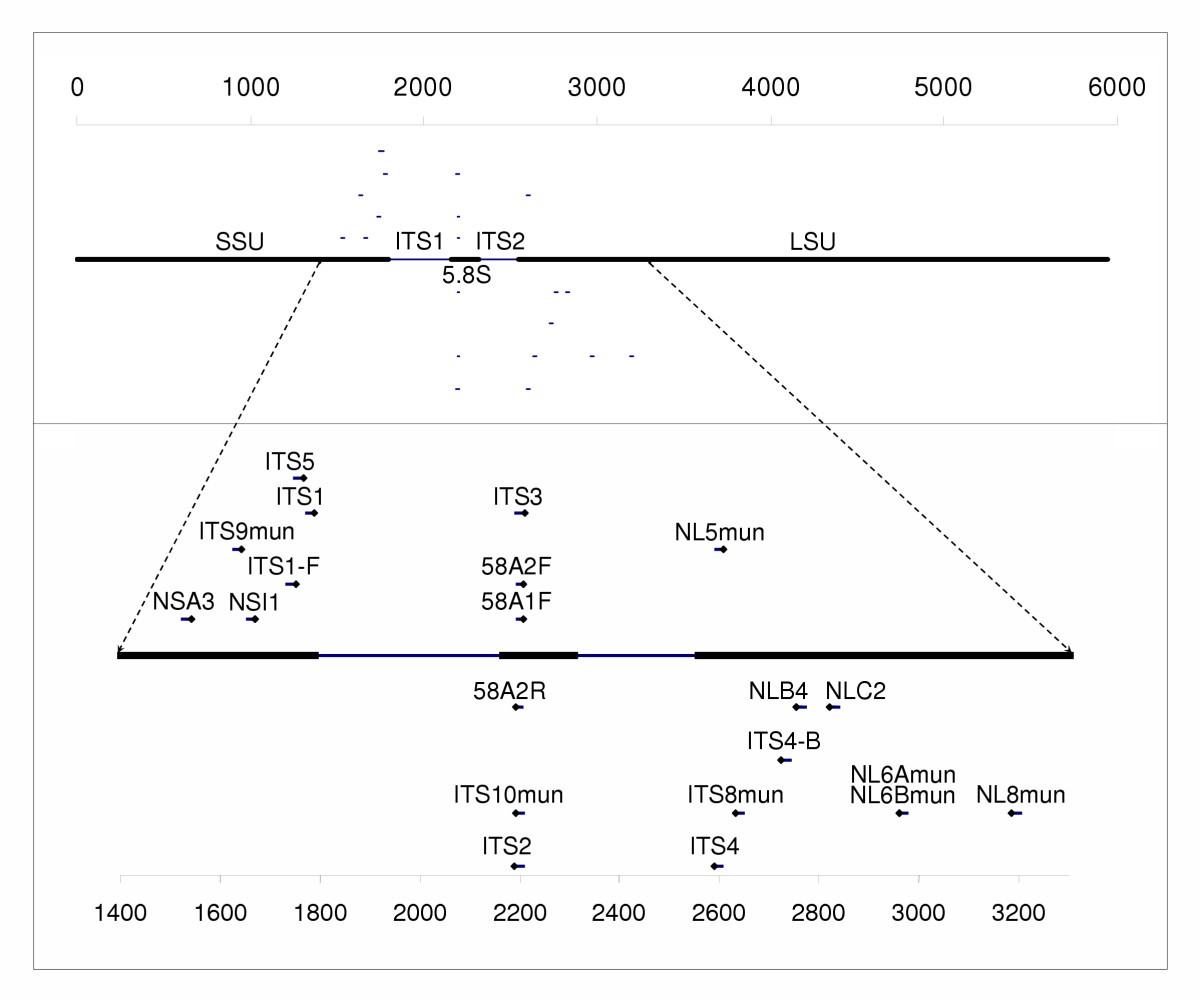
Fungal-specific PCR primers developed for analysis of the ITS region of environmental DNA extracts | BMC Microbiology | Full Text
![Comparison of the performance of ITS1 and ITS2 as barcodes in amplicon-based sequencing of bioaerosols [PeerJ] Comparison of the performance of ITS1 and ITS2 as barcodes in amplicon-based sequencing of bioaerosols [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/8523/1/fig-5-2x.jpg)
Comparison of the performance of ITS1 and ITS2 as barcodes in amplicon-based sequencing of bioaerosols [PeerJ]
![Internal Transcribed spacer (ITS) region primers ITS1 and ITS 4 [8]... | Download Scientific Diagram Internal Transcribed spacer (ITS) region primers ITS1 and ITS 4 [8]... | Download Scientific Diagram](https://www.researchgate.net/publication/264669072/figure/fig1/AS:614269146112029@1523464585883/Internal-Transcribed-spacer-ITS-region-primers-ITS1-and-ITS-4-8-used-for.png)
Internal Transcribed spacer (ITS) region primers ITS1 and ITS 4 [8]... | Download Scientific Diagram

IJMS | Free Full-Text | Intra-Genomic Internal Transcribed Spacer Region Sequence Heterogeneity and Molecular Diagnosis in Clinical Microbiology

Representative gel electrophoresis of the ITS1 and ITS4 PCR products.... | Download Scientific Diagram
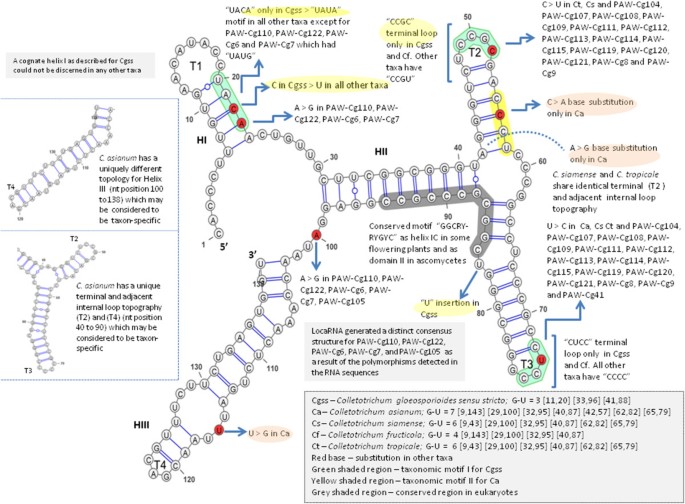
ITS1, 5.8S and ITS2 secondary structure modelling for intra-specific differentiation among species of the Colletotrichum gloeosporioides sensu lato species complex | SpringerPlus | Full Text
![Comparison of the performance of ITS1 and ITS2 as barcodes in amplicon-based sequencing of bioaerosols [PeerJ] Comparison of the performance of ITS1 and ITS2 as barcodes in amplicon-based sequencing of bioaerosols [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/8523/1/fig-4-full.png)
Comparison of the performance of ITS1 and ITS2 as barcodes in amplicon-based sequencing of bioaerosols [PeerJ]

ITS1: a DNA barcode better than ITS2 in eukaryotes? - Wang - 2015 - Molecular Ecology Resources - Wiley Online Library
![Comparison of the performance of ITS1 and ITS2 as barcodes in amplicon-based sequencing of bioaerosols [PeerJ] Comparison of the performance of ITS1 and ITS2 as barcodes in amplicon-based sequencing of bioaerosols [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/8523/1/fig-6-2x.jpg)
Comparison of the performance of ITS1 and ITS2 as barcodes in amplicon-based sequencing of bioaerosols [PeerJ]